Introduction — The Hidden Vulnerabilities of Bacteria
Antimicrobial drugs succeed because bacteria, despite their remarkable adaptability, rely on biological systems that differ in important ways from those of human cells. These differences create opportunities for selective intervention. An antimicrobial compound that interferes with a process unique to bacteria can cripple a pathogen while leaving human tissues largely unaffected. This fundamental principle underlies the field of antibacterial therapy and explains why antibiotics have remained such powerful tools in medicine for more than 80 years.
A single microbial cell operates an intricate network of molecular machinery that maintains its structure, produces proteins, replicates its genetic material, and generates the metabolic intermediates needed for growth and division. Most antibiotics work by interfering with an activity essential to the survival of the pathogenic bacteria.
Viewed conceptually, antimicrobial therapy resembles sabotage inside a factory. A drug might disable the workers building the outer wall, jam the assembly line that produces proteins, damage the copying machines that reproduce genetic instructions, or block the supply chains that deliver essential biochemical components. When any of these core systems fails, bacterial growth slows or stops, and in many cases, the cell dies.
Although hundreds of antibiotic molecules have been discovered, most act through a limited number of underlying strategies. These mechanisms fall into several broad categories: inhibition of cell wall construction, disruption of protein synthesis, interference with DNA or RNA production, blockade of key metabolic pathways, and damage to the bacterial membrane itself. Understanding these mechanisms provides a useful framework for exploring both the strengths of existing antimicrobial drugs and the challenges posed by the evolution of resistance.
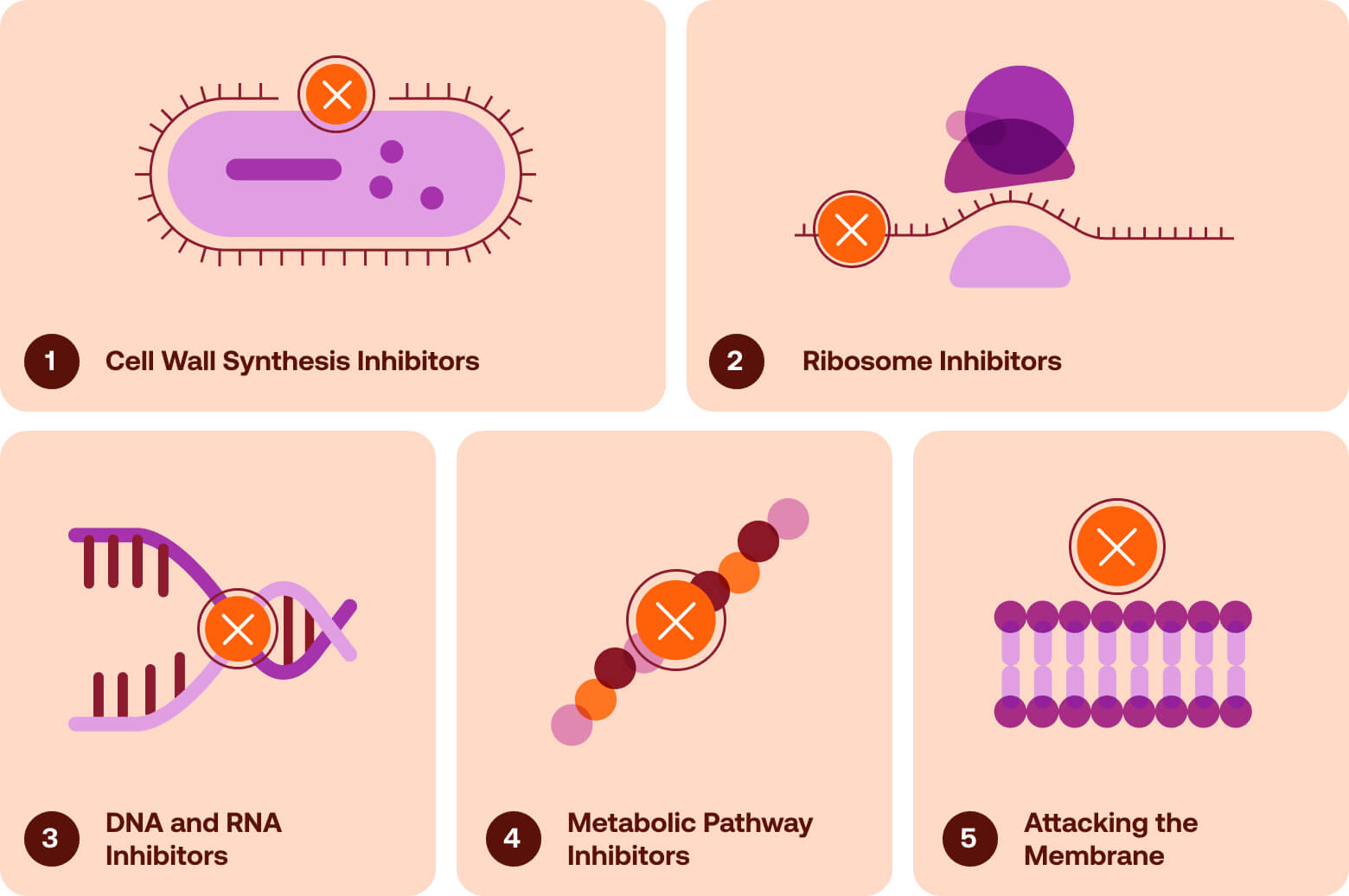
Breaking the Walls: Cell Wall Synthesis Inhibitors
For many bacteria, survival depends on the integrity of the cell wall, a rigid mesh-like shell surrounding the membrane that protects the cell from osmotic pressure. Because the concentration of dissolved molecules inside the cell is often higher than in the surrounding environment, water tends to flow inward. The cell wall provides the mechanical strength needed to prevent swelling and rupture, allowing bacteria to maintain their shape and structural stability.
The wall itself is built primarily from peptidoglycan, a lattice of sugar chains connected by short peptide bridges. These cross-linked strands create a resilient network analogous to reinforced concrete. Because human cells lack this structure entirely, the enzymes responsible for assembling peptidoglycan are attractive, highly selective targets for antimicrobial drugs.
Among the most important cell wall inhibitors are the β-lactam antibiotics, a family that includes penicillins, cephalosporins, carbapenems, and related compounds. These drugs act by interfering with the final stages of peptidoglycan assembly. During normal cell wall construction, penicillin-binding proteins (PBPs) connect peptide strands to create the cross-linked framework that gives the wall its strength. β-lactams bind covalently to PBPs, inhibiting their transpeptidase activity and preventing cross-linking of peptidoglycan strands. As a result, newly synthesized cell wall material becomes structurally weak and unstable.1
The consequences for the bacterium are often catastrophic. As the cell continues to grow and divide, defects accumulate in the wall. Eventually, the weakened structure can no longer withstand internal pressure, leading to rupture of the cell membrane and the organism’s death. For this reason, β-lactam antibiotics are typically bactericidal rather than merely bacteriostatic. Their ability to kill bacteria directly and relatively low toxicity to human cells have made them one of the most widely used and clinically successful antibiotic classes in medical history.1
Despite their effectiveness, β-lactams are also vulnerable to resistance. Many bacteria produce β-lactamases, enzymes that chemically degrade the antibiotic before it can reach its target. Others acquire altered PBPs that bind β-lactams less effectively while still carrying out cell wall synthesis. These adaptations allow pathogens to preserve the essential cross-linking process even in the presence of the drug.2
A second group of cell wall inhibitors takes a different approach to the same biological system. Glycopeptide antibiotics, such as vancomycin, do not disable the enzymes responsible for assembling the wall. Instead, they bind directly to the molecular building blocks used to construct it. During peptidoglycan synthesis, precursor molecules containing a terminal d-alanine–d-alanine (d-Ala–d-Ala) motif are incorporated into the growing structure. Vancomycin binds tightly to this motif, preventing precursors from being incorporated into the cell wall lattice.3
This strategy effectively removes the raw materials needed for construction. While the enzymes responsible for assembly remain functional, they cannot access the substrates required to extend the wall. As a result, bacterial growth stalls and the structural integrity of the cell envelope gradually fails. Glycopeptides have therefore become particularly important for treating infections caused by Gram-positive organisms, including pathogens that are resistant to many other antibiotics.
Resistance to glycopeptides arises through a different mechanism than resistance to β-lactams. Instead of destroying the drug or altering an enzyme target, some bacteria modify the structure of the cell wall precursor itself. In certain resistant strains, the d-Ala–d-Ala motif is replaced with d-Ala–d-Lac. This subtle chemical change reduces vancomycin binding dramatically while still allowing peptidoglycan assembly to proceed.4
Sabotaging the Protein Factory: Ribosome Inhibitors
If the cell wall provides structural protection, proteins provide function. Enzymes catalyze metabolic reactions, structural proteins maintain cellular organization, and regulatory proteins coordinate responses to environmental changes. Nearly every activity required for bacterial survival depends on the continuous production of new proteins. Interrupting this process quickly slows growth and, in some cases, kills the cell outright.
Protein production takes place in ribosomes, molecular complexes that translate genetic instructions encoded in messenger RNA (mRNA) into chains of amino acids. Bacterial ribosomes differ structurally from their human counterparts, allowing drugs to interfere with microbial protein synthesis without disrupting the same process in human cells. For this reason, the ribosome has been one of the most productive targets in antimicrobial therapy.
Several antibiotic families interfere with protein synthesis, each exploiting a different vulnerability in the ribosome’s operation. Some distort the translation process itself, while others physically block the movement of molecular components needed for protein assembly.
Aminoglycoside drugs bind to the 30S ribosomal subunit and interfere with the decoding of mRNA. Under normal circumstances, the ribosome reads genetic instructions three nucleotides at a time, matching each codon to the corresponding transfer RNA (tRNA) that carries a specific amino acid. Aminoglycosides disrupt this proofreading process, causing the ribosome to misread the message. The resulting proteins contain incorrect amino acids and often fail to fold properly or perform their intended functions.5,6
The accumulation of defective proteins can damage cellular processes in several ways. Some proteins lose their enzymatic activity, while others become unstable or interfere with essential pathways. In severe cases, the errors introduced during translation lead to the production of toxic proteins that compromise membrane integrity or other critical systems. Because of these cascading effects, aminoglycosides often display rapid bactericidal activity rather than merely slowing bacterial growth.6
Bacteria can resist aminoglycosides through several mechanisms, but one of the most common involves enzymatic modification of the drug itself. Certain bacterial enzymes chemically alter the antibiotic molecule through processes such as acetylation, phosphorylation, or adenylation. These modifications reduce the drug’s ability to bind the ribosome, allowing protein synthesis to continue even in the presence of the antibiotic.
Macrolide antibiotics take a different approach to disrupting protein production. Instead of causing decoding errors, these drugs physically obstruct part of the ribosome’s architecture. Macrolides bind to the large, 50S ribosomal subunit within a channel known as the nascent peptide exit tunnel. As new proteins are synthesized, the growing amino acid chain normally passes through this tunnel and emerges from the ribosome. When a macrolide occupies this space, the passage becomes partially blocked, preventing the newly forming peptide from extending properly.7
This interference stalls the ribosome during translation, halting protein synthesis before the molecule is complete. Because macrolides act at a defined structural site within the ribosome, relatively small changes to that site can confer resistance. A common mechanism involves methylation of the ribosomal RNA that lines the exit tunnel. This chemical modification alters the shape of the binding pocket, reducing macrolides’ ability to bind to the ribosome.8
A third group of ribosome-targeting antibiotics, the tetracyclines, disrupts protein synthesis at an earlier step in the process. These drugs bind to the 30S subunit at a region known as the A-site, where incoming tRNA molecules normally deliver amino acids to the ribosome. By occupying this position, tetracyclines prevent aminoacyl-tRNA from entering the ribosome and participating in translation.9
Without a steady supply of tRNA molecules bearing amino acids, translation stalls, and the bacterium is unable to produce the proteins required for metabolism, replication, and structural maintenance. Tetracyclines therefore act primarily as bacteriostatic agents, slowing bacterial growth and allowing the immune system to clear the infection.
Resistance to tetracyclines often arises through one of two strategies. Efflux pumps embedded in the bacterial membrane can actively export the drug from the cell, reducing its intracellular concentration. Alternatively, specialized ribosomal protection proteins can dislodge tetracyclines from the ribosome, restoring the ability of tRNA molecules to bind and resume translation.10
Disrupting the Blueprint: DNA and RNA Inhibitors
Bacteria depend on accurate transmission of genetic information, which requires DNA replication to copy the genome and transcription to produce messenger RNA (mRNA) that guides protein synthesis. Several antibiotic classes exploit this vulnerability by interfering with the enzymes responsible for these processes.
Fluoroquinolones target the machinery that allows bacterial chromosomes to be replicated and organized. As a circular DNA molecule is copied, it becomes tightly twisted and tangled. Specialized enzymes known as DNA gyrase and topoisomerase IV relieve this mechanical strain by cutting the DNA, allowing it to unwind, and then resealing the strands. Fluoroquinolones bind to these enzymes and stabilize an intermediate stage in this process in which the DNA has been cleaved but not yet rejoined.11
This stabilization traps the enzyme–DNA complex in a dysfunctional state. Instead of relieving tension in the chromosome, the enzyme remains locked onto broken DNA strands, preventing replication from proceeding. As these stalled complexes accumulate, double-strand DNA breaks can form, leading to lethal damage within the bacterial genome.12 Because these enzymes play such central roles in chromosome replication and organization, inhibiting them can halt bacterial growth across a wide range of species. Fluoroquinolones thus exhibit broad-spectrum antibacterial activity and have been widely used to treat infections of the respiratory and urinary tracts and other systems.
Resistance to fluoroquinolones most often arises through mutations in the genes encoding DNA gyrase or topoisomerase IV. These mutations subtly alter the enzyme’s structure, causing the antibiotic to bind less effectively while the enzyme continues to perform its normal function. In addition, some bacteria employ efflux pumps that actively export fluoroquinolones from the cell, reducing the concentration of the drug available to interact with the target enzymes.
While fluoroquinolones disrupt DNA replication, rifamycins interfere with transcription by binding to the β-subunit of bacterial RNA polymerase. Once attached, the drug blocks the extension of newly formed RNA chains, effectively preventing the transcription process from progressing beyond its earliest steps.13
Without the ability to produce RNA transcripts, bacteria cannot generate the mRNA molecules required for protein synthesis. The entire chain of information flow — from DNA to RNA to protein — breaks down. Rifamycins therefore exert a powerful antibacterial effect and have become central components of treatment regimens for diseases such as tuberculosis.
Resistance to rifamycins typically arises from mutations in the rpoB gene, which encodes the RNA polymerase β-subunit. These mutations alter the antibiotic’s binding site, reducing its ability to interact with the enzyme while leaving the transcription machinery functional.14 This mechanism illustrates a recurring theme in antimicrobial resistance: small genetic changes in a drug target can allow bacteria to evade inhibition while preserving essential cellular processes.
Starving the Cell: Metabolic Pathway Inhibitors
Not all antibiotics act by damaging major cellular structures or halting the production of proteins and nucleic acids. Some instead target the metabolic pathways that supply bacteria with the molecules required to sustain growth. These pathways function as biochemical supply chains, producing the building blocks needed for DNA synthesis, amino acid metabolism, and other essential cellular processes. Interrupting one of these pathways deprives the bacterium of key compounds, slowing or stopping its ability to replicate.
One of the best-known examples involves the folate biosynthesis pathway. Folate molecules play a central role in the synthesis of nucleotides. Without a reliable supply of folate, bacteria cannot produce the genetic material needed for cell division. Unlike humans, who obtain folate from dietary sources, many bacteria must synthesize it internally, providing a selective opportunity for antimicrobial therapy.
Two closely related drug classes exploit this pathway: sulfonamides and trimethoprim. Sulfonamides interfere with an early step in folate production by inhibiting dihydropteroate synthase, which normally helps assemble the precursor molecules that ultimately lead to the formation of dihydrofolate, an intermediate in the folate pathway. When sulfonamides block this step, the bacterium cannot produce sufficient folate to support DNA synthesis.15
Trimethoprim acts further downstream in the same pathway. It inhibits dihydrofolate reductase, the enzyme that converts dihydrofolate to tetrahydrofolate, the active form required for nucleotide synthesis and other metabolic reactions.16 When this conversion is blocked, the folate cycle stalls, and the cell is unable to produce the nucleotides needed for replication.
Because these drugs target sequential steps in the same pathway, they are often used together in combination therapy. The paired inhibition creates a metabolic bottleneck that is far more difficult for bacteria to bypass than either drug alone would be. This synergy enables the combination to achieve greater antibacterial activity while reducing the likelihood of resistance arising from simple alterations at a single enzyme target.
Attacking the Membrane: Last-Line Agents
Beyond the cell wall lies another critical defensive structure: the bacterial membrane. This lipid-based barrier encloses the cytoplasm and regulates the movement of molecules into and out of the cell. In gram-negative bacteria, the membrane system is even more complex, consisting of both an inner membrane and an outer membrane that contains lipopolysaccharide (LPS). Together, these layers create a formidable protective envelope that shields the bacterium from environmental stress and many antimicrobial agents. Because membrane integrity is essential for survival, drugs that damage this structure can rapidly compromise bacterial viability.
Polymyxins exploit this vulnerability. These molecules are cyclic peptides with a strong affinity for lipid A, a component of LPS that lines the outer membrane of Gram-negative bacteria. Under normal conditions, lipid A molecules are stabilized by interactions with divalent cations that help maintain the tight packing of the membrane surface. Polymyxins displace these stabilizing ions and bind directly to lipid A, disrupting membrane organization.17
This disruption weakens the outer membrane and increases its permeability. As the membrane structure deteriorates, essential cellular contents can leak out, and toxic molecules can enter. The resulting damage interferes with multiple cellular processes and ultimately leads to bacterial death. Because the mechanism involves direct destabilization of the membrane rather than inhibition of a specific enzyme, polymyxins can remain effective against gram-negative pathogens that resist many other antibiotics.
For this reason, polymyxins are often reserved as last-line treatments for severe infections caused by multidrug-resistant organisms, including difficult pathogens, such as Acinetobacter baumannii, Pseudomonas aeruginosa, and certain strains of Klebsiella pneumoniae. Their ability to breach the outer membrane makes them particularly valuable in situations where other drugs cannot reach their intracellular targets.
Resistance to polymyxins typically arises through modifications of the bacterial outer membrane itself. Some bacteria alter the chemical structure of lipid A, reducing polymyxins’ ability to bind effectively. These modifications alter the membrane surface charge, weakening the electrostatic interactions that normally attract the antibiotic molecule.18 Although such resistance mechanisms are still relatively uncommon compared with resistance to older antibiotic classes, their emergence highlights the continuing evolutionary pressure faced by even the most potent antimicrobial agents.
Why Resistance Is Inevitable
The remarkable adaptability of bacteria constantly challenges the effectiveness of antimicrobial drugs. Microbial populations reproduce quickly, and each generation introduces opportunities for genetic variation. Within these vast populations, even rare mutations can provide a survival advantage when antibiotics are present. Over time, natural selection favors organisms that can withstand the drug’s effects, allowing resistant strains to expand while susceptible ones disappear.
Mutation is not the only path through which resistance emerges. Bacteria can also acquire resistance genes from other organisms through a process known as horizontal gene transfer. Unlike vertical inheritance, in which genetic changes pass from parent cells to their descendants, horizontal gene transfer allows bacteria to exchange genetic material directly across species and even across genera. Resistance genes are frequently carried on plasmids — small, circular DNA molecules that replicate independently of the bacterial chromosome — or on mobile genetic elements such as transposons that can move between different DNA molecules. When these elements carry genes encoding β-lactamases, efflux pumps, or modified drug targets, they can spread resistance traits rapidly through microbial populations. In environments where antibiotics are present, this genetic exchange accelerates the emergence of multidrug-resistant strains and enables resistance mechanisms discovered in one species to disseminate across many others rapidly.
Several well-established mechanisms enable bacteria to evade antimicrobial activity. One common strategy involves producing enzymes that chemically destroy or modify the antibiotic before it reaches its target. β-lactamases provide a classic example: these enzymes break the β-lactam ring structure of penicillins and related drugs, rendering them inactive. Similar enzymatic processes can alter other antibiotic classes, preventing the drug from binding effectively to its intended target.19
Another route to resistance involves changes to the molecular structures that antibiotics normally attack. If a mutation alters the shape of a target protein or enzyme, the drug may no longer bind tightly enough to inhibit its function. The bacterium retains the ability to perform the essential cellular process while avoiding the drug’s inhibitory effects. Mutations in penicillin-binding proteins, ribosomal RNA, or DNA gyrase illustrate how small genetic changes can produce large shifts in antibiotic susceptibility.
Bacteria can also prevent antibiotics from reaching their targets in the first place. Efflux pumps embedded in the cell membrane actively transport certain drugs out of the cell, lowering intracellular concentrations to levels that no longer interfere with bacterial processes. Other resistance mechanisms involve changes in membrane permeability, particularly in gram-negative bacteria whose outer membranes act as selective barriers. By altering membrane channels or reducing the uptake of specific compounds, bacteria can limit the entry of antibiotics into the cell. 20
These adaptations illustrate an important reality of antimicrobial therapy: resistance is not simply the result of poor drug design or clinical misuse, although both can accelerate its spread. Instead, resistance emerges naturally whenever selective pressure favors organisms capable of surviving antimicrobial exposure. Understanding the biological mechanisms underlying resistance is therefore essential for developing new therapeutic strategies and preserving the effectiveness of existing drugs.

Expanding the Toolbox: Emerging Mechanisms
Although most existing antibiotics operate through a relatively small set of biological strategies, ongoing research continues to identify new ways to exploit bacterial vulnerabilities. Rather than focusing solely on incremental improvements to existing drug classes, many current discovery efforts aim to identify previously untapped targets or novel ways of interacting with known ones. These approaches do not replace established antibiotics, but they broaden the therapeutic toolbox and may help address pathogens that have developed resistance to older mechanisms.
One promising direction involves targeting the molecular precursors used to construct the bacterial cell wall. Teixobactin, a compound discovered through screening of previously uncultured soil microorganisms, illustrates this strategy. Instead of binding enzymes that assemble the wall, teixobactin attaches directly to lipid II and lipid III, essential building blocks of peptidoglycan synthesis. By sequestering these precursors, the drug prevents their incorporation into the growing cell wall, halting construction before it can begin.21,22 Because these lipid intermediates are highly conserved components of bacterial physiology, this mechanism may narrow the range of mutations that could readily confer resistance.
Another area of investigation focuses on the complex envelope surrounding Gram-negative bacteria. These organisms possess an outer membrane that provides an additional protective barrier and contributes to their resistance to many antibiotics. Several experimental compounds disrupt the machinery responsible for assembling this outer layer. Darobactin, for example, targets BamA, the central component of the β-barrel assembly machinery (BAM) complex that inserts proteins into the outer membrane. By interfering with this process, the drug prevents the insertion of outer-membrane proteins required for membrane integrity.23 Other candidates act on the systems that transport lipopolysaccharide molecules to the bacterial surface. Zosurabalpin has been shown to interfere with the LptB2FGC transporter that extracts lipopolysaccharide from the inner membrane during outer-membrane assembly.24 At the same time, murepavadin targets the related LptD component involved in lipopolysaccharide transport in P. aeruginosa.25 These strategies highlight how envelope biogenesis itself can serve as a drug target.
Novel mechanisms are also emerging even for targets that have been exploited for decades. Odilorhabdins show that a well-known structure, such as the ribosome, can still yield new opportunities for intervention. These peptides bind to the bacterial ribosome at a site not targeted by existing antibiotics, thereby altering translation accuracy and interfering with protein synthesis through a distinct mechanism.26,27
Finally, some experimental drugs target metabolic pathways that antibiotics have not traditionally exploited. Compounds that inhibit FabI, an enzyme involved in bacterial fatty acid biosynthesis, illustrate this approach. Because fatty acid production is essential for maintaining the bacterial membrane and other cellular structures, inhibiting this pathway can impair bacterial growth while potentially sparing beneficial microbes that rely on different metabolic systems.28
Conclusion — The Continuing Evolution of Antimicrobial Strategy
Despite the wide range of antibiotic molecules discovered over the past century, most successful antimicrobial drugs exploit a relatively small set of biological vulnerabilities. These strategies have proven remarkably effective because they target processes that bacteria cannot easily dispense with while leaving human cells largely unaffected.
At the same time, the evolutionary capacity of bacteria ensures that resistance will remain an ongoing challenge. Mutations that alter drug targets, enzymes that inactivate antibiotics, and cellular systems that reduce drug accumulation can all diminish the effectiveness of existing therapies. As these adaptations spread through microbial populations, antibiotics that once worked reliably may become less effective or require combination therapy to remain useful. Understanding the molecular basis of both antibiotic action and resistance, therefore, remains central to maintaining the effectiveness of antimicrobial treatments.
The continuing search for new antibiotics reflects this dynamic relationship between microbial evolution and drug discovery. Emerging compounds demonstrate that bacterial physiology still contains untapped vulnerabilities. Some experimental agents target components of cell-envelope assembly that were previously overlooked, while others bind familiar targets, such as the ribosome, in entirely new ways. Still others interfere with metabolic processes that earlier antibiotic classes have rarely exploited.
Antimicrobial strategy is not limited to the mechanisms discovered in the early decades of antibiotic research. Instead, it continues to evolve as researchers gain deeper insight into bacterial biology. Expanding the range of drug targets, improving stewardship of existing antibiotics, and developing therapies that reduce the likelihood of resistance will all play roles in sustaining the effectiveness of antimicrobial medicine in the years ahead.