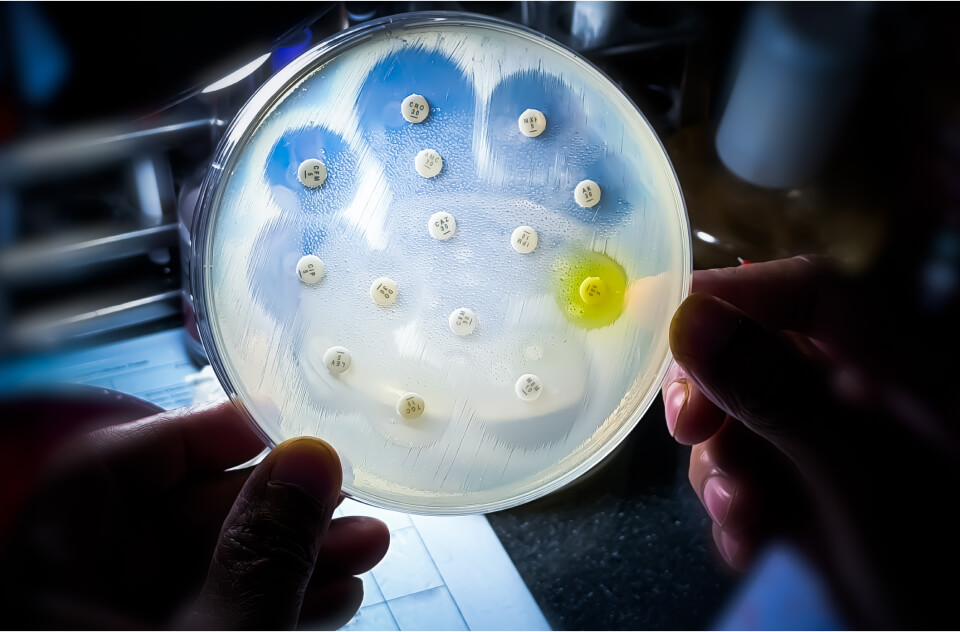
Antibiotic Resistance Continues to Outpace Drug Development
The number of bacterial pathogens capable of resisting multiple antibiotics continues to rise, including resistance to some recently introduced drugs.1,2 This growing challenge is reflected in the World Health Organization’s (WHO) priority pathogen list, which identifies organisms for which urgently needed new therapies remain scarce.3 At the top of this list are several gram-negative bacteria — Acinetobacter baumannii, Pseudomonas aeruginosa, and members of the Enterobacterales family — along with major gram-positive threats, such as Staphylococcus aureus, Enterococcus spp., and Streptococcus pneumoniae. Collectively, these organisms account for a large share of difficult-to-treat infections in both hospital and community settings.
The problem extends beyond classical genetic resistance. Many pathogenic bacteria acquire mutations that allow them to survive exposure to antibiotic molecules that would previously have been lethal. Other populations display antibiotic tolerance, remaining viable (though not actively proliferating) during transient drug exposure. Still others form persistent subpopulations that enter dormant states capable of surviving even very high antibiotic concentrations.4 These multiple survival strategies allow pathogens to withstand treatment, recover after therapy ends, and ultimately contribute to the continued expansion of antimicrobial resistance.
As a result, even as new antibiotics enter the market, many of them remain vulnerable to the same biological defenses that undermine older drugs. Addressing antimicrobial resistance will therefore require approaches that extend beyond traditional antibiotic mechanisms and draw on a broader therapeutic toolkit.
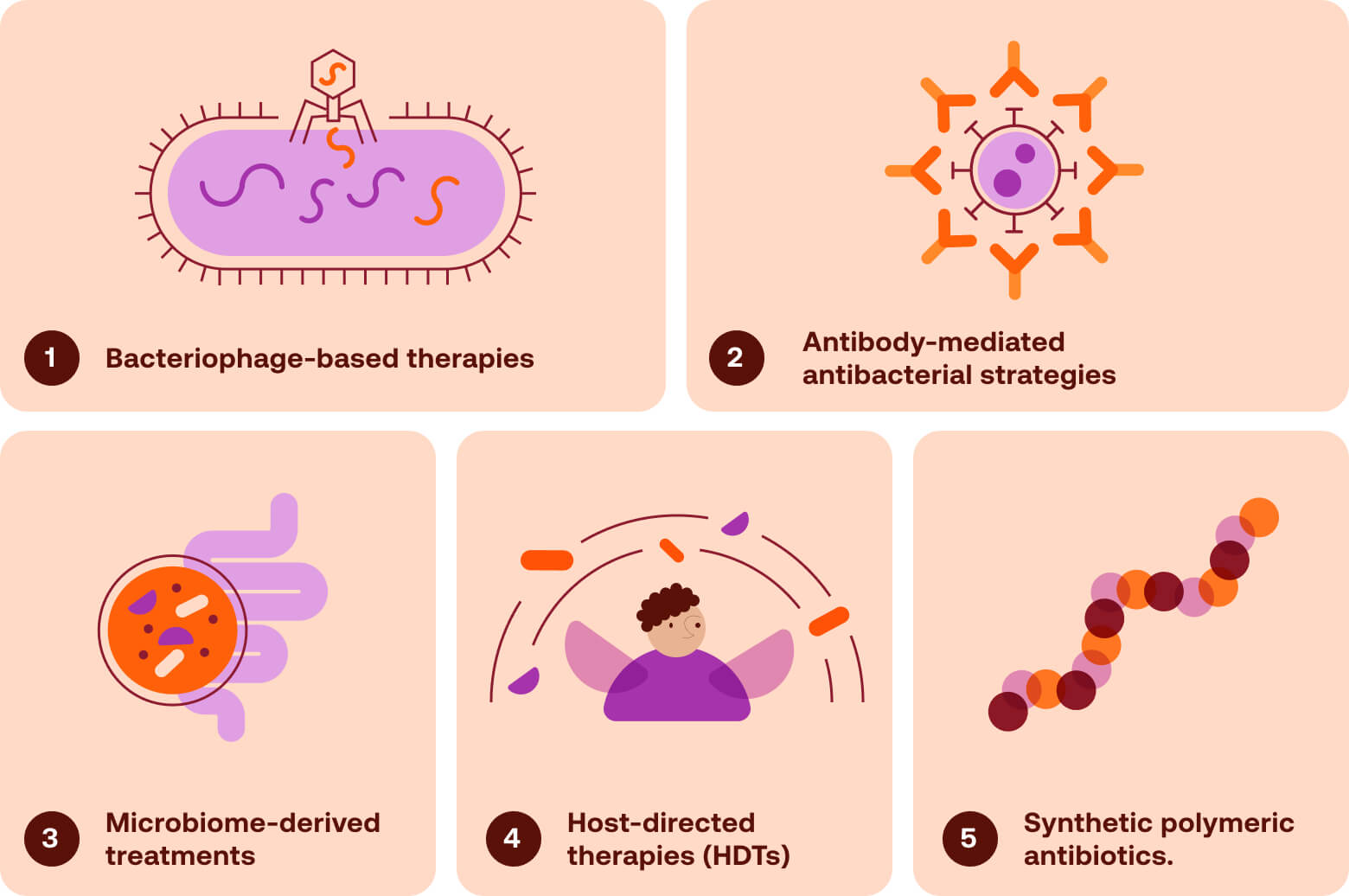
Moving Beyond the Classical Antibiotic Paradigm
Analyses published in early 2020 by both the WHO and the Pew Charitable Trusts reached a sobering conclusion: none of the antibiotics currently available worldwide can reliably address many clinically significant drug-resistant bacterial infections.4 While some new antibiotics have been introduced in recent years, most represent incremental improvements rather than fundamentally new solutions. Between the publication of the WHO priority pathogen list in 2017 and late summer 2025, the U.S. Food and Drug Administration (FDA) approved 16 antibiotics. Yet, nearly all were from previously established antimicrobial classes.1 As a result, many remain vulnerable to resistance mechanisms that bacteria have already evolved.
Traditional antibiotics operate through a relatively limited set of biological mechanisms. Most act by inhibiting bacterial cell wall synthesis, disrupting cell membrane function, or interfering with protein synthesis by binding to ribosomal subunits.5 Bacteria can evade these effects through several well-established strategies, including reducing drug uptake, accelerating drug metabolism, enzymatically inactivating antibiotics, or modifying the molecular targets to which the drugs bind.4,6 Once these mechanisms emerge and spread, new drugs that rely on the same underlying modes of action often face the same fate.
Expanding the antimicrobial arsenal will therefore require therapies that operate outside the traditional antibiotic paradigm and bypass these entrenched resistance pathways. A growing body of research is exploring alternative therapeutic modalities that target bacteria through fundamentally different mechanisms. Prominent examples include bacteriophage-based therapies, antibody-mediated antibacterial strategies, microbiome-derived treatments, host-directed therapies that strengthen immune defenses, and synthetic polymeric antibiotics.
Harnessing Bacteriophages to Combat Drug-Resistant Bacteria
Bacteriophage therapy is among the most conceptually distinct alternatives to traditional antibiotics. Bacteriophages, a class of viruses that infect and kill bacteria, target specific bacterial species and replicate within them, creating a self-amplifying antimicrobial effect. Each phage attaches to a bacterial cell and introduces its genetic material, initiating one of two possible biological cycles. In the lytic cycle, the phage takes over the host cell’s machinery to produce new phage particles, ultimately causing the bacterial cell to rupture and release additional viruses. In the lysogenic cycle, phage DNA integrates into the bacterial chromosome or persists in the cytoplasm as an episome, becoming a prophage that is replicated as the bacterium divides and can later transition into the lytic cycle under environmental stress.7
Because bacteriophages infect bacteria rather than human cells, they do not damage host tissues and generally do not disrupt the broader human microbiome like broad-spectrum antibiotics.8 Their high specificity also allows targeted elimination of pathogenic bacteria while leaving beneficial microbial communities largely intact. Another advantage lies in their evolutionary adaptability: although bacteria can develop resistance to individual phages, phages themselves can evolve in response to bacterial defenses, creating a dynamic biological arms race that differs fundamentally from static chemical antibiotics.
The therapeutic use of bacteriophages is not a new concept. Phage therapy was first explored more than a century ago and continues to be used in parts of Eastern Europe as a treatment option when conventional antibiotics fail.7 Modern research has expanded on this foundation, demonstrating that phage therapy often performs best when multiple phages are combined into phage cocktails, which broaden the range of bacterial targets and reduce the likelihood that resistance will emerge.
Recent preclinical studies involving pathogens, such as P. aeruginosa, A. baumannii, and Klebsiella pneumoniae, have shown promising results.9 Early clinical experience, largely derived from compassionate-use cases, suggests that combining phages with conventional antibiotics may further enhance effectiveness in patients with multidrug-resistant infections. These phage–antibiotic combinations (PACs) have been reported to enhance bacterial killing, suppress the emergence of resistance, and, in some cases, restore susceptibility to antibiotics that had previously lost efficacy.10
Despite these advances, important practical challenges remain. Many phage therapies must be tailored to the specific bacterial strain causing a patient’s infection, requiring rapid identification of the pathogen and testing candidate phages against it.9,10 Standardized clinical workflows for evaluating patient isolates are still limited, and the time required for strain identification and phage matching can restrict the use of personalized phage therapy in rapidly progressing infections.
To address these limitations, researchers are developing off-the-shelf phage cocktails designed to cover broader bacterial populations. One recent study employed a screening strategy to identify “complementarity groups” of phages targeting different bacterial receptors and to examine their interactions with specific antibiotic classes. The resulting phage–antibiotic combinations demonstrated high efficacy against P. aeruginosa and S. aureus individually and when used together against mixed bacterial cultures.11
Although these findings highlight the potential of phage therapy as a powerful addition to the antimicrobial toolkit, the field still lacks large, systematically designed clinical trials. Existing studies involve diverse phage–antibiotic combinations administered through varying routes, dosing schedules, and treatment durations, making it difficult to draw definitive conclusions about optimal therapeutic strategies.10 Continued clinical investigation will be necessary to determine how phage-based therapies can be integrated most effectively into the treatment of drug-resistant infections.
Importantly, phage therapy illustrates how expanding beyond conventional antibiotics can introduce entirely different biological strategies for controlling bacterial infections: approaches that exploit viral replication and evolutionary dynamics rather than relying solely on static chemical inhibition.
Antibody Therapies: Precision Targeting of Bacterial Pathogens
Monoclonal antibodies (mAbs) represent another strategy for expanding the antimicrobial toolkit beyond conventional antibiotics. Traditional small-molecule antibiotics, while effective in many settings, often act broadly against bacteria and can disrupt beneficial microbial communities as well as pathogens. They may also persist in the environment and exert selective pressure that accelerates the development of resistance. These limitations have prompted growing interest in antibody-based approaches that target bacterial pathogens with far greater precision.12,13
Antibodies can be engineered to recognize highly specific bacterial antigens, allowing them to attack pathogenic organisms while minimizing collateral damage to the surrounding microbiome. Their pharmacokinetic properties also offer practical advantages: many antibodies exhibit relatively long half-lives in circulation, which can reduce dosing frequency and potentially improve treatment adherence. Because they are produced via cell culture rather than chemical synthesis and degrade naturally in biological environments, antibody therapeutics may also offer environmental advantages over traditional antibiotics.
Unlike small molecules, which typically disrupt bacterial physiology directly, antimicrobial antibodies act through a range of immune-mediated mechanisms. These include neutralizing bacterial toxins, blocking receptor-mediated bacterial adhesion to host tissues, interfering with biofilm formation, promoting immune clearance through antibody-mediated complement killing, and enhancing opsonophagocytosis (antibody tagging of pathogens that facilitates their engulfment by phagocytic immune cells). Antibodies can also bind outer-surface proteins associated with antibiotic resistance, thereby weakening bacterial defenses. By mobilizing multiple immune mechanisms rather than relying on a single biochemical target, antibody-based therapies may reduce the likelihood that bacteria will rapidly develop resistance.12,13
The versatility of antibody engineering further expands their therapeutic potential. Small molecule antibiotics can be conjugated to antibodies to create antibody–antibiotic conjugates, enabling targeted delivery of antibacterial payloads directly to infected cells while limiting off-target bacterial killing. Bispecific antibodies can simultaneously bind two different bacterial antigens or epitopes, creating a coordinated multipoint attack. Researchers are also exploring alternative antibody formats, including nanobodies and single-chain antibody fragments (scFvs), which may offer improved tissue penetration or manufacturing advantages.
Several antibody therapies targeting bacterial pathogens have already reached the market. Examples include raxibacumab (GlaxoSmithKline) and obiltoxaximab (Elusys Therapeutics), which neutralize Bacillus anthracis toxins in bioterrorism preparedness applications, and bezlotoxumab (Zinplava, Merck & Co.), which helps prevent recurrence of Clostridioides difficile infection in high-risk patients.12
Despite these advances, clinical development of antibacterial antibodies has proven challenging. Only a limited number of candidates have progressed into late-stage trials, and many programs have stalled during phase I or phase II studies.13 Early efforts often focused on fully humanized antibodies targeting pathogens, such as S. aureus, P. aeruginosa, or Clostridium botulinum. Several factors contributed to their limited clinical success. Bacterial infections frequently involve multiple pathogens, antigenic variability can reduce antibody effectiveness, and conventional monoclonal antibodies often rely on a single mechanism of action. For example, antibodies that neutralize bacterial toxins may limit tissue damage after infection has begun but may not directly eliminate the underlying bacteria.
Newer strategies aim to address these limitations by expanding the functional capabilities of antibody therapeutics. Antibody–antibiotic conjugates combine the targeting precision of antibodies with the bactericidal activity of conventional antibiotics, while bispecific antibodies can simultaneously target multiple bacterial antigens. Researchers are also investigating antibody cocktails that include several antibodies targeting different targets, potentially broadening their activity against diverse pathogens and reducing the likelihood of resistance.13
Microbiome Therapeutics: Restoring the Body’s Natural Defenses
The human microbiome plays a central role in maintaining resistance to infection, and disruption of this complex microbial ecosystem can significantly increase vulnerability to pathogenic bacteria. Such disruption can lead to dysbiosis, defined as an imbalance in the composition and functional capacity of microbial communities.14 Dysbiosis has been associated with numerous disease states,15 and the gut microbiota also plays an important role in drug metabolism and host immune regulation.14 When the composition or activity of the microbiome changes, susceptibility to infection can increase as protective microbial functions are weakened.16
These microbial communities influence host biology through multiple mechanisms. Alterations in microbiome composition can affect the production of antimicrobial peptides, influence epigenetic regulation and gene expression patterns, modify enzymatic activity through microbiome-derived metabolites, and shape inflammatory and cellular immune responses. Changes in microbial populations can also alter pH levels, influence nutrient availability for pathogens, and modify drug pharmacokinetics, including the activity and effectiveness of antibiotics.16 Through these interconnected pathways, microbiome disruptions can create conditions that allow pathogenic bacteria to establish infections more easily.
Microbiome-based therapeutics aim to restore or reshape these microbial ecosystems rather than directly killing pathogens. By promoting beneficial microbial populations, these therapies can help reestablish colonization resistance and reduce susceptibility to infection without the broad-spectrum bacterial killing often associated with conventional antibiotics.4 Several classes of microbiome-directed therapies are under development, including prebiotics, which serve as substrates for beneficial microbes; probiotics, which introduce live microorganisms with defined biological functions; synbiotics, which combine prebiotics and probiotics; and postbiotics, which comprise microbial metabolites or cellular components that influence microbiome activity.14
Probiotics in particular have attracted significant interest as researchers and biotechnology companies explore the use of defined microbial consortia — often referred to as live biotherapeutic products (LBPs) — to treat infectious and inflammatory diseases.17 Candidate organisms under investigation include Akkermansia muciniphila, Faecalibacterium prausnitzii, Veillonella, Ruminococcus, Christensenella minuta, and Bacteroides fragilis.14 Advances in genetic engineering are also enabling the design of modified microbial strains with enhanced targeting capabilities or therapeutic functions.16
One of the clearest demonstrations of microbiome-based therapy involves the treatment of recurrent or multidrug-resistant C. difficile infection (CDI). In these cases, fecal microbiota transplantation (FMT) from healthy donors has proven highly effective in restoring microbiome diversity and preventing recolonization by C. difficile spores. A donor-derived microbiome therapy, Rebyota (Ferring Pharmaceuticals), has received FDA approval for this indication.14 More recently, oral capsule formulations containing lyophilized microbiota have been developed, offering a simpler and less invasive treatment option. VOWST (Seres Therapeutics) is the first FDA-approved product of this type. Researchers are also evaluating microbiome transplantation approaches for other multidrug-resistant gastrointestinal infections and recurrent bacterial vaginal infections.14
Additional strategies under investigation include microbiome-based vaccines and combination approaches in which microbiome therapeutics are administered alongside conventional antibiotics to enhance treatment outcomes.16 By reinforcing the body’s natural microbial defenses, these approaches illustrate another pathway for expanding antimicrobial strategies beyond traditional antibiotic mechanisms.
Turning the Host into the Antimicrobial Target
Another strategy for addressing antimicrobial resistance focuses on strengthening the host rather than directly attacking bacterial pathogens. Host-directed therapies (HDTs) seek to enhance innate or adaptive immune responses or modulate cellular pathways that pathogens rely on for survival and replication.18,19 Because these approaches target host processes rather than bacterial structures, pathogens may find it more difficult to evolve resistance. HDTs interfere with intracellular and intercellular pathways that bacteria exploit during infection, thereby disrupting the conditions required for pathogen persistence.18,20
Several host-directed strategies have been explored in the treatment of chronic infectious diseases, such as tuberculosis. Many of these therapies focus on regulating inflammatory signaling pathways. Agents targeting cytokine activity, including antibodies directed against interleukin-6 or tumor necrosis factor, have been investigated alongside compounds such as interferons, antioxidants like N-acetylcysteine, vitamins including vitamin D, and anti-inflammatory drugs, such as statins and cyclooxygenase-2 inhibitors.18,20 By modulating immune signaling, these therapies aim to restore effective host responses while limiting excessive inflammation that can damage tissues.
Other HDTs work by stimulating innate antimicrobial mechanisms within host cells. Research has explored compounds that increase production of antimicrobial peptides (AMPs), reactive oxygen species (ROS), and nitric oxide (NO), as well as therapies that induce autophagy, a cellular process that degrades intracellular pathogens.21 These molecules and pathways form part of the body’s natural antimicrobial defenses and contribute to the barrier against microbial invasion. Many multidrug-resistant pathogens suppress these protective mechanisms during infection. Therapies that stimulate their production can therefore help restore the host’s ability to eliminate invading bacteria. In some cases, these mechanisms may act synergistically when activated together.
Epigenetic regulators provide another route for activating antimicrobial defenses. Histone deacetylase inhibitors (HDACis), such as butyrate and other short-chain fatty acids (SCFAs), can increase the production of antimicrobial peptides while strengthening epithelial barrier function by promoting tighter cell junctions.21 However, direct administration of butyrate can trigger undesirable immunogenic reactions in the gastrointestinal tract, limiting its clinical utility. Alternative molecules such as phenylbutyrate have demonstrated promising activity in animal models. Vitamin D has also attracted interest because it promotes autophagy and can enhance antimicrobial activity when combined with other host-directed therapies.
Several classes of drugs already approved for other diseases may also be repurposed as host-directed antimicrobial therapies. Immunomodulatory drugs, certain cellular and gene therapies, and antibody-based treatments originally developed for cancer are now being evaluated for infectious diseases.19 Current research focuses heavily on chronic infections, such as tuberculosis, hepatitis B and C, and HIV-1, but similar strategies may prove applicable to a broader range of bacterial pathogens.
Despite their promise, host-directed therapies present important challenges. Because these interventions alter host cellular pathways, they carry the risk of unintended toxicities.19 Their effectiveness may also depend on careful timing relative to infection stage and disease progression. Successful HDT development, therefore, requires identifying host pathways that pathogens depend upon but that can be safely modulated without disrupting essential physiological functions.
Programmable Polymers for Combating Drug-Resistant Bacteria
Synthetic antimicrobial polymers (SAPs) represent another emerging strategy for combating drug-resistant infections. Unlike conventional antibiotics that typically interfere with specific intracellular targets, many SAPs act directly on bacterial membranes. Their antimicrobial activity derives in part from their relatively large molecular size and in part from the ability to engineer their chemical structure by selecting different polymer backbones and functional substituents.22 Because most polymers cannot readily penetrate bacterial cells, they instead interact with the negatively charged outer surfaces of both gram-positive and gram-negative bacteria. Positively charged polymers, including poly(quaternary ammonium) compounds, polyethylenimine, and polyamines, can bind to these surfaces and disrupt membrane integrity, leading to bacterial cell death through mechanisms that are inherently difficult for pathogens to evade through conventional resistance pathways.22
Researchers have explored a wide range of SAP architectures. Some polymers exhibit intrinsic antimicrobial activity, while others carry attached bioactive functional groups or act as delivery vehicles for antibacterial agents.22 Most SAPs are cationic, but anionic and amphiphilic polymers containing both positive and negative charges have also been investigated. Functional groups incorporated into polymer backbones often include cationic, hydrophobic, π-electron–containing, amphiphilic, and metal-binding moieties, each of which can influence antimicrobial activity. In addition, stimuli-responsive polymers are being designed to activate under specific environmental conditions, such as changes in pH or temperature, or exposure to oxidative environments or enzymes, enabling more targeted antimicrobial effects.
Many SAPs have already been applied as antimicrobial coatings for medical devices and implants to reduce contamination and infection risk.22 Increasingly, however, these materials are being investigated as systemic antimicrobial therapies. Experimental polymers have demonstrated activity against pathogens such as Escherichia coli and Salmonella, including multidrug-resistant strains.
A closely related class of materials — synthetic antimicrobial peptides (AMPs) — has also attracted significant interest. Natural AMPs are key components of innate immune defenses, and synthetic variants can be engineered to disrupt bacterial membranes while also influencing immune and inflammatory responses.23 The development of clinical AMP candidates has proven challenging due to issues such as toxicity, limited in vivo stability, and manufacturing complexity. Advances in computational design are helping address these limitations. Deep learning approaches are now being used to identify peptide sequences with improved antimicrobial activity and safety profiles.
Machine learning is also accelerating the broader design of antimicrobial polymers. In one recent example, researchers identified three poly(β-amino ester) polymers with strong activity against drug-resistant bacteria, low toxicity, and the ability to self-assemble into stable nanoparticles (SANPs).23 When combined with penicillin G, these polymers restored susceptibility in drug-resistant gram-positive bacteria, illustrating how polymeric materials can enhance the effectiveness of existing antibiotics.
Other approaches combine polymer chemistry with traditional antibiotics to overcome specific resistance mechanisms. One self-assembling nanocomplex composed of a guanidinium-functionalized cationic polymer and the anionic prodrug colistin methane sulfonate was developed to target bacteria carrying plasmid-borne mobilized colistin resistance (mcr) genes.24 By delivering both the antibiotic and the antimicrobial polymer simultaneously, this system was able to reverse colistin resistance and effectively kill mcr-positive strains of multidrug-resistant E. coli and K. pneumoniae.
Polyacrylamide-based copolymers represent another promising platform. These materials disrupt bacterial membranes while maintaining selectivity for bacterial cells over mammalian cells and have demonstrated efficacy against several multidrug-resistant pathogens.25 When administered alongside conventional antibiotics, these SAPs can also enhance antibiotic activity against resistant bacteria.
These polyacrylamide polymers offer several practical advantages, including commercial availability, chemical stability, and relatively straightforward synthesis.25 Effective candidates typically incorporate a cationic monomer to support membrane binding, a hydrophilic monomer to maintain water solubility, and a smaller fraction of hydrophobic monomer designed to promote membrane disruption. Leading candidates have demonstrated activity against pathogens such as Staphylococcus aureus, E. coli, K. pneumoniae ATCC 13884 (gram-negative), and Enterococcus faecium ATCC 35667 (gram-positive). Notably, four of these polymers showed activity against all four pathogens.
Not all antimicrobial polymers are large macromolecules. Researchers have also developed shorter, biodegradable oligomeric polymers capable of penetrating bacterial membranes. One example involves poly(imidazolium ester) polymers that cross bacterial membranes and interact directly with nucleic acids.26 In vitro studies showed that these polymers bind DNA through electrostatic and hydrogen-bond interactions to form biomolecular condensates. This phase separation disrupts transcription and translation processes and may also interfere with RNA synthesis. In preclinical animal models, the leading candidate safely treated systemic, intramuscular, and pulmonary infections caused by drug-resistant bacteria and fungi, including S. aureus, S. epidermidis, E. coli, K. pneumoniae, and Candida albicans, at relatively low doses.
By enabling antimicrobial activity through engineered membrane disruption, programmable nanostructures, and novel intracellular mechanisms, polymer-based therapeutics demonstrate how materials science and molecular engineering can broaden the range of strategies for combating antimicrobial resistance.
Rethinking Antibiotic Development for the AMR Era
Scientific innovation alone will not resolve the antimicrobial resistance crisis if economic and development incentives remain misaligned. One of the most persistent barriers to progress is the limited investment in antibiotic development by major pharmaceutical companies. According to one recent analysis, only six large biopharma companies currently maintain small antibiotic development programs, while several biotechnology firms focused on antibiotic innovation have entered bankruptcy.27 This contraction reflects a difficult economic reality: new antibiotics are often reserved for the most severe or resistant infections to preserve their effectiveness, which significantly limits the revenue they can generate.
These economic constraints have prompted some researchers to question whether the traditional strategy of developing highly targeted antibiotics for individual resistant pathogens is sustainable. Treatments designed to address specific multidrug-resistant organisms often serve relatively small patient populations, and the growing diversity of resistant bacteria would require developing many separate drugs to address each threat.27 As a result, some investigators have proposed a shift toward “pathogen-general” antimicrobial approaches that could work across a wider range of bacterial species.
One proposed solution involves the development of general potentiators — agents that enhance the effectiveness of existing antibiotics against both gram-positive and gram-negative bacteria, including strains that have already developed resistance.27 Rather than replacing current antibiotics, these therapies would restore or amplify their activity, potentially extending the useful life span of established treatments. Ideally, such agents would combine broad applicability with practical advantages in manufacturability, cost, safety, and clinical deployment.
Many of the emerging modalities discussed earlier, including phage therapy, host-directed therapies, antibody-based antibacterial strategies, microbiome-directed treatments, and polymeric antibiotics, could contribute to this type of pathogen-general strategy. By either enhancing host defenses, targeting bacterial virulence mechanisms, reshaping microbial ecosystems, or sensitizing bacteria to existing drugs, these approaches may complement traditional antibiotics and expand the range of tools available for managing resistant infections.
Antimicrobial resistance will not be solved by simply discovering more versions of the same antibiotics that bacteria have already evolved resistance to. Instead, progress will likely come from a broader therapeutic ecosystem that combines multiple complementary strategies. Phage therapies, antibody-based antibacterial agents, microbiome-directed treatments, host-directed approaches, and engineered antimicrobial polymers each operate through fundamentally different mechanisms that can either bypass traditional resistance pathways or restore the effectiveness of existing drugs. As these modalities mature and enter clinical development, they may work alongside conventional antibiotics rather than replace them outright. Expanding the antimicrobial toolkit in this way could provide the flexibility needed to address the diverse and evolving challenges posed by multidrug-resistant pathogens.